We Will Summarize The Results That Transformer Has Brought To Medical Imaging.
3 main points
✔️ Comprehensive review of medical imaging and Transformer
✔️ Reviews have been conducted on detection, classification, reconstruction, synthesis, registration, clinical reporting, and other tasks and the latest information can be found on the authors' GitHub.
✔️ Transformer has penetrated all areas of medical imaging and is progressing too rapidly. To cope with this, it is desirable to organize workshops at conferences, publish special issues in journals, and quickly disseminate related research to the community.
Transformers in Medical Imaging: A Survey
written by Fahad Shamshad, Salman Khan, Syed Waqas Zamir, Muhammad Haris Khan, Munawar Hayat, Fahad Shahbaz Khan, Huazhu Fu
(Submitted on 24 Jan 2022)
Comments: Published on arxiv.
Subjects: Image and Video Processing (eess.IV); Computer Vision and Pattern Recognition (cs.CV)
code:

The images used in this article are from the paper, the introductory slides, or were created based on them.
Abstract
Transformers have been successfully used in natural language and have also been applied to computer vision (CV), where convolutional neural networks (CNNs) are dominant, but interest in Transformers is of increasing interest, and so is the medical field. This is also true in the medical field, and this paper provides a comprehensive review of how Transformers are superior to CNNs with local receptive fields in capturing the local context in the whole image.
Introduction
CNN has had a significant impact on the field of medical imaging. Significant improvements have been reported in many areas such as radiography, endoscopy, CT, MRI, mammography, PET, and ultrasound. However, CNNs, by their nature, are not good at capturing the meaning of local images about the whole image or how distant objects in the image are related to each other.
On the other hand, a new architecture has been explored in the CV domain, inspired by the attention mechanism (note: we write "attention" in the text). Attention-based Transformer enables us to learn long-range dependencies as effective feature representations, which is an attractive method to solve the above problems. Recent work has shown that standard CNNs can be completely replaced by Transformers, giving rise to Vision Transformers (ViTs). Since their inception, ViTs have achieved remarkable classification, detection, segmentation, and colorization. It has also been shown that the prediction error of ViTs is similar to that of humans, which has also sparked interest in the medical domain.
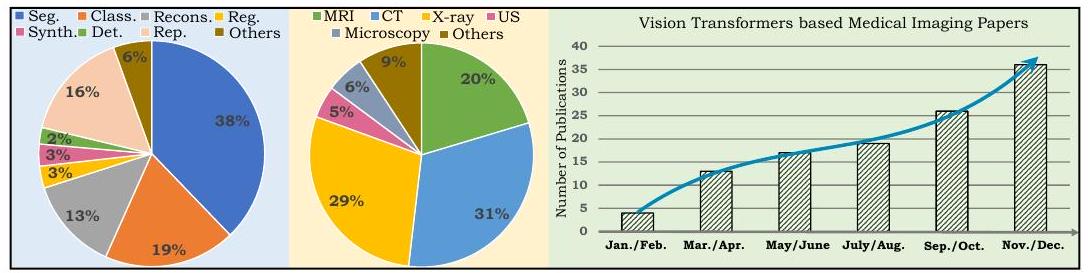
The pie chart on the left shows the breakdown of papers covered by this review, where Seg indicates segmentation, Class indicates classification, Recons indicates reconstruction, Reg indicates registration, Synth indicates synthesis, Det indicates detection, Rep indicates reporting, and the US indicates ultrasound. The chart on the right shows the number of papers published in 2021 that apply ViT to medical imaging, showing a rapid increase.
Background
Handcrafted Mathematical Models
Traditional algorithms in medical imaging tasks are based on expert-designed mathematical models (hand-crafted approaches). This approach was refined in the direction of being less computationally intensive and applicable to a larger number of tissues. The result was models that were interpretable and robust with mathematical underpinnings and widely used in the medical domain. The approach was different from today's deep learning and did not require large data sets or annotations.
CNN-based methods
CNN is a method for automatically learning discriminative features from large datasets. It has shown excellent performance in medical imaging and has become an integral part of modern AI-based medical systems. However, the performance-dependent nature of CNNs (essentially) depends on the size of the dataset, which limits their adaptation to the medical domain. In addition, CNN inference results are generally difficult to interpret and often function in a black box manner.
Transformer-based methods
The transformer was reported in the natural language field as an attention-driven block. Attention is a NN that aggregates information from an entire input sequence. Since the introduction of attention, several models have rewritten the best performance. It is now the first choice for model selection.
ViT is a model built by connecting multiple Transformer layers to capture the global context of the input image.
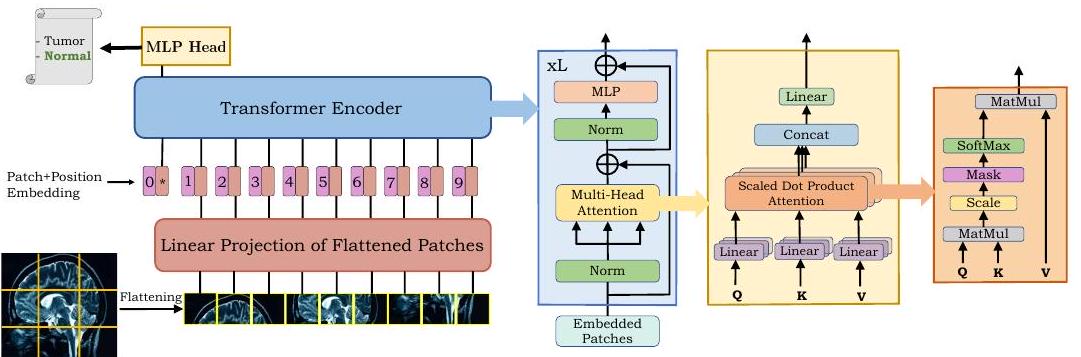
As shown in the figure above, ViT divides the image into tiles (patches) and arranges them in a sequence. This sequence is encoded by Transformer to produce the final output.
Self-Attention
The success of Transformer is attributed to the self-attention mechanism (self-attention, SA), which models long-range dependence. The key idea of self-attention is to determine the relative importance of a single token in the whole sequence.
Multi-Head Self-Attention
Multi-head self-attention (MHSA) consists of multiple blocks (heads) to model the complex dependencies of the input sequence. while MHSA allows learning complex contexts, attention is computationally expensive, which is a limitation in medical imaging where pixel counts are often high. This is a limitation in medical imaging, where the pixel count tends to be high. Therefore, improved and more efficient attention has been proposed for medical image processing.
Segmentation Field
We begin with an overview of medical image segmentation. Segmentation is important for computer-aided diagnosis (CAD), image-guided surgery, and treatment planning. For example, organs have large spatial extents, so it is necessary to model the relationship between distant pixels. This is where Transformer's global context modeling capabilities come in handy. Conversely, the background of an ultrasound image is noisy and scattered. By modeling the background, it is possible to recognize the areas that are necessary for diagnosis.
Organ segmentation
two-dimensional segmentation
Wang et al. applied ViT to skin lesions.
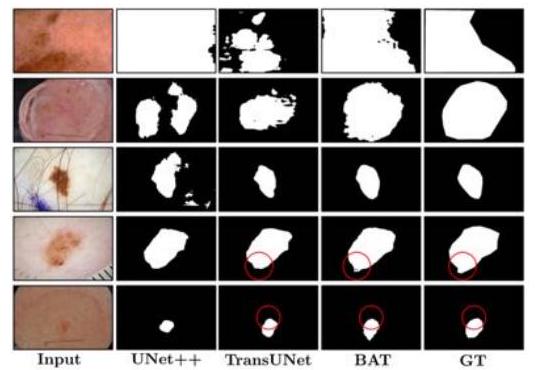
They proposed the Boundary-Aware Transformer (BAT) and succeeded in extracting a clear boundary between melanoma and normal skin.
Other ViT-based methods have been proposed for dental roots, vascular vessels, kidney tumors, cells, and tissues.
3D Segmentation
Wang et al. utilized a Transformer to model the spatial features of brain tumors. Their proposed TransBTS used a 3D CNN to extract local 3D space and a Transformer to encode more global features. Compared to simple CNNs, TransBTS is noteworthy for its effectiveness and for eliminating the need for pre-training on large datasets, which is required for traditional ViT.
Zhu et al. performed 3D segmentation in breast cancer. The proposed method, RAT-Net, outperformed previously proposed CNN and ViT-based methods.
multiorgan segmentation
Multi-organ segmentation is the segmentation of multiple organs at once. It is a more difficult task than single-organ segmentation due to imbalances between classes, differences in organ size, shape, and contrast.
Pure Transformer
Pure Transformer-based architectures consist of only a ViT layer. Since segmentation requires two perspectives, global and local, hybrid CNN-Transformer architectures are often proposed. sets and showed its effectiveness on three benchmark sets: brain cortex, spleen, and hippocampus.
hybrid architecture
Hybrid architectures have been proposed, using Transformers to model the global context and CNNs for accurate segmentation. The first proposed method is TransUNet, which uses 12 Transformer layers.
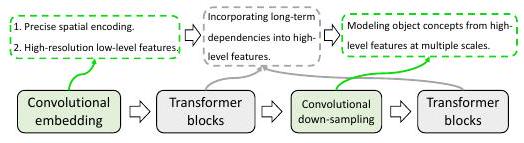
As shown above, Transformer and CNN are connected in the hybrid architecture so that they can show their advantages.
Segmentation Discussion
ViT-based segmentation has received significant attention with more than 50 reports per year. In this domain, simply converting CNN-based methods to Transformer-based ones has resulted in improved performance in most cases. However, the computational cost is high, which is a limitation in the field of medical imaging.
Many of the ViT models also use pre-training models in ImageNet. However, it has been pointed out that there is a modality gap between natural images and medical images, which makes them unsuitable for pre-training. Recent studies on image modalities used in pre-training have shown that models pre-trained on CT do not perform satisfactorily when applied to MRI. This modality difference is an issue for future research.
medical image classification
A tumor is an abnormal growth of tissue in the body, which can be benign or malignant. Their diagnosis is very important for treatment planning and contributes significantly to patient survival.
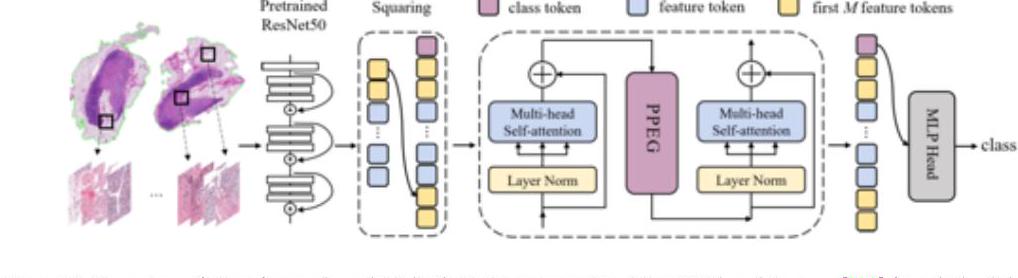
In the above TransMIL, a patch of whole slide images (WSI) is embedded in the ResNet-50 feature space. TransMIL achieves SOTA in three tissues: mammary, lung, and kidney.
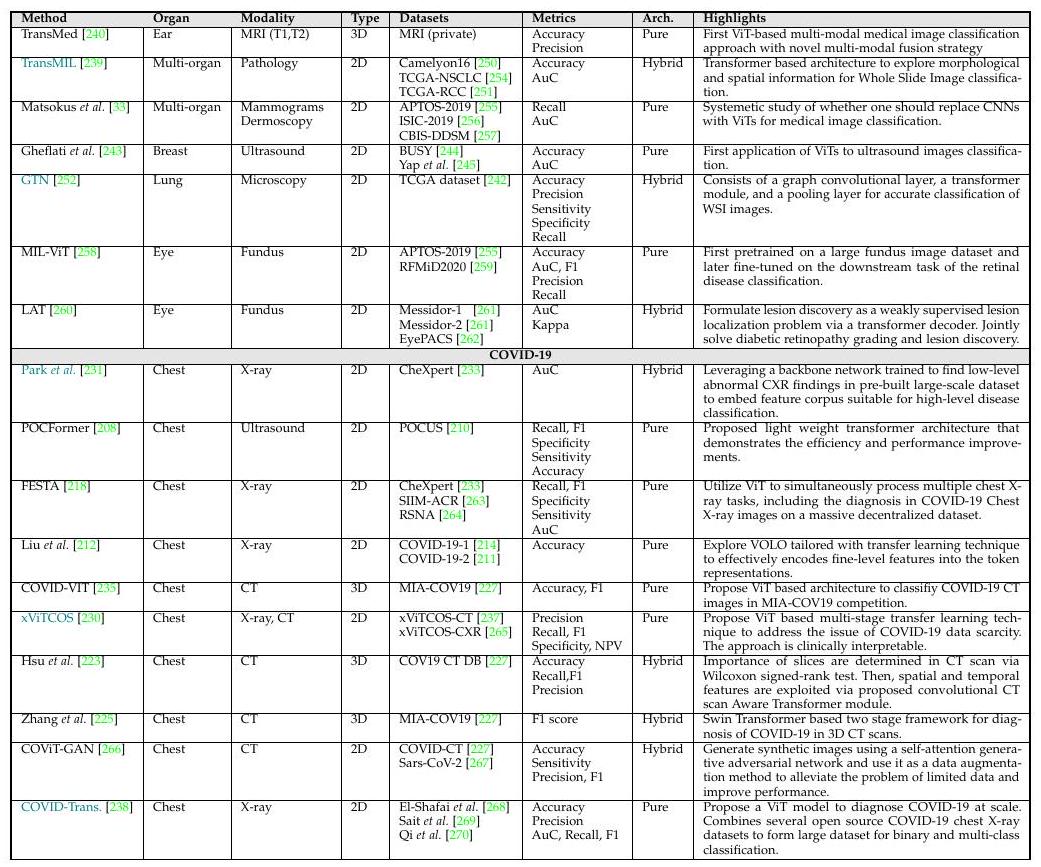
Above is an overview of ViT-based medical image classification. Methods in blue are models that also show Grad-CAM interpretability.
Classification Discussion
In the classification task, it was found that most of the studies used the original ViT and made no changes (plug-and-play). Therefore, it is expected that the addition of domain-specific architectures, design of loss functions, etc. will lead to the proposal of effective ViT-based models in the future.
Object detection in medical images
Object detection in medical imaging refers to the localization of regions of interest (ROI), for example, finding lung nodules in a chest radiograph. Transformers have recently been used to achieve even higher performance. Many of these approaches are based on the Detecting Transformer (DETR) framework.
The COTR proposed by Shen et al. has been applied to polyp detection and lymph node detection on T2-weighted MR images and has shown better performance than DETR.
Discussion on Object Detection
Transformer-based object detection models are few compared to segmentation and class classification studies, in contrast to the early development of object detection in CNN-based models. Therefore, it is expected that more Transformer-based research will be reported shortly.
Reconstruction in medical images
The goal of the reconstruction field is to obtain a clean image from a degraded input (note: the literal translation of reconstruction is reconstruction, which has a broad meaning. (Note: The literal translation of reconstruction is to reconstruct, which is a broad term that can be thought of as a generic term for restoration or correction).
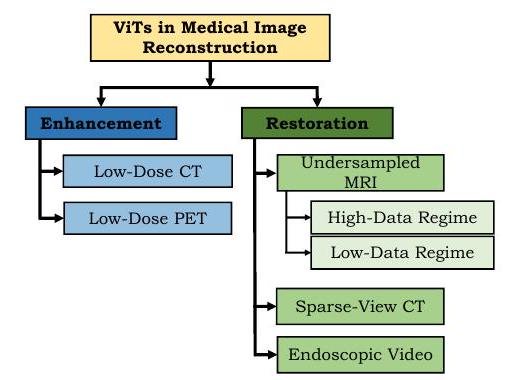
ViT has made remarkable progress in image enhancement. Low-dose radiographs are blurred, but by enhancing the image quality, it is possible to maintain or improve diagnostic accuracy while reducing radiation doses.
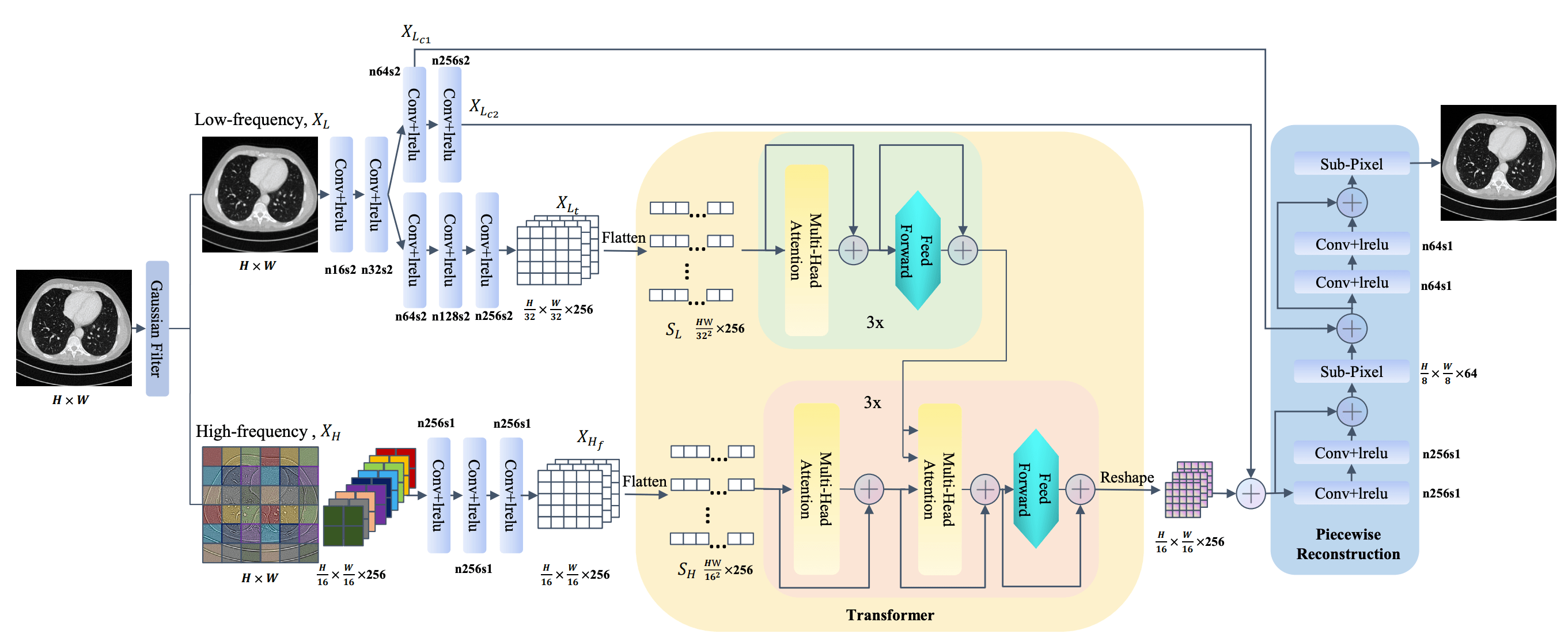
As shown above, Zhan et al. proposed TransCT, which effectively enhances low-dose CT images.
Similarly, reducing the number of measurements per cross-section in MR can also reduce motion artifacts.
Discussion in Reconstruction
We surveyed 12 papers in this review, many of which were for CT images. In addition, the approaches are generic, so we expect that future work will be more task-specific in terms of loss functions and architectures, which will lead to further performance improvements.
medical image synthesis
In the application of ViT to medical image synthesis, many approaches incorporate adversarial loss. In this field, there are two main types of composting: intra-modality (intra-modality) and inter-modality (inter-modality) (note: modality refers to the type of imaging, such as CT, MR, etc.).
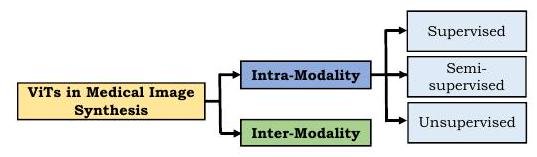
For synthesis in the same modality, supervised, semi-supervised, and unsupervised learning methods have been proposed. In the case of supervised learning, a pair of teacher images and its corresponding target image is required, which is costly not only for medical images. In the case of medical images, it is extra limited by the difficulty of collection and the cost of annotation. However, Zhang et al. proposed PTNet to synthesize an infant's brain MRI and demonstrated that the model is qualitatively and quantitatively better than the previously proposed pix2pix and pix2pixHD. Furthermore, PTNet is not only high performance but also remarkable for its moderate execution time of about 30 seconds per image.
The inter-modality approach acquires images of different modalities and outputs a target image. inter-modality methods only use supervised learning because it is a modality conversion task.
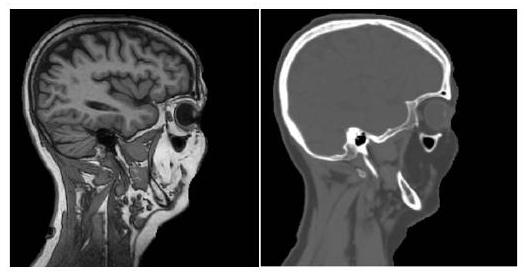
The inter-modality method converts the MRI (left) into a CT image (right) as shown above. The results are excellent, with completely different appearances.
Discussion on Medical Image Synthesis
In general, clinical decision-making involves images from multiple modalities that are complementary to each other (note: CT looks at hard tissue such as bones and teeth, while MRI looks at soft tissue such as muscle, brain, and tumors). However, from a cost perspective, multiple modality images are not always available; Transformer-based image synthesis has been found to produce more realistic images than GAN-based methods, effectively avoiding these problems.
Registration of medical images
Registration refers to the alignment between two images. For example, CT and MR images of the same patient may not overlap exactly due to the position of the images or subtle body twists.
TransMorph proposed by Chen et al. reveals the correspondence between still images and videos. To capture the semantic correspondence between the input image and video, it is encoded by the Swin Transformer and the subsequent decoder is used to infer the displacement.
Discussion in Registration
At this stage, Transformer is not widely used in the registration area, and it is difficult to summarize. However, as Transformer-based methods in general imaging are developing, we expect that they will develop in the medical field in the future as well.
Reporting
In recent years, the research area of automatic report generation from medical images has been developed. However, the task of creating a reading report or adding captions is very challenging because the reports written by humans (radiologists) themselves are varied and of varying length.
The task requires two things: first, that the language is expressed in a human-readable manner, and second, that the content is medically accurate.
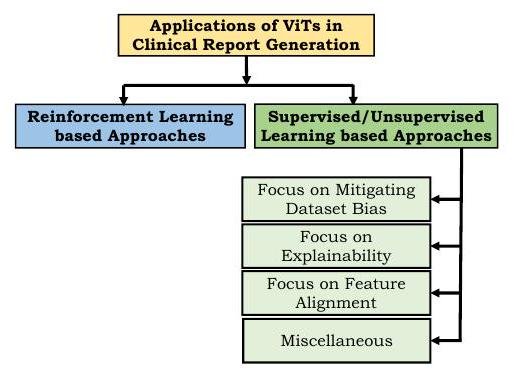
Reinforcement learning based approaches use the medical terms used, human ratings, etc. as rewards; one of the first attempts using Transformer was the Reinforced-Transformer for Medical Image Captioning (RTMIC) proposed by Xiong et al. (RTMIC) proposed by Xiong et al. It consists of a DenseNet to identify ROIs from medical images and a Transformer-based encoder to extract visual features, and the decoder part generates captions.
Supervised and unsupervised learning approaches use differentiable loss functions to train models for diagnostic writing. However, many medical reports contain far more sentences describing normality than abnormality, resulting in a dataset bias. To reduce this bias, Srinivasan proposed a hierarchical classification approach using a Transformer as a decoder. This architecture consists of an abnormality detection part to classify normal or abnormal images, a tag generation part for images, and a sentence generation part from image features and tags, and its effectiveness was demonstrated on chest radiographs.
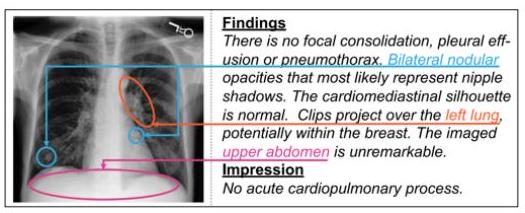
To improve confidence in the model, an attempt is also made to indicate which parts of the image are of interest for the resulting text.
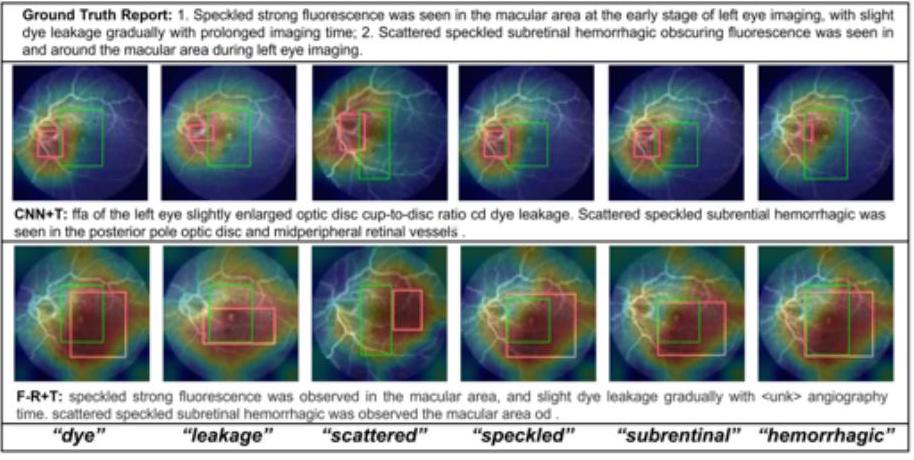
As shown in the figure above, we can show the correspondence between the text and the focus area in the CNN+Transformer and Faster-RCNN+Transformer models, and we can see from which part of the image the words dyeing and leakage are recalled. the words "dye" and "leakage".
Report Writing Discussion
We have reviewed ViT so far, but the focus has been on Transformer as a powerful language model when it comes to reporting. Despite the impact of Transformer's phenomenal uptake since 2017, the number of reports on clinical reporting is low.
Transformer-based models use natural language generation (NLG) metrics such as CIDEr and BLEU. However, NLG metrics do not evaluate clinical expressions well. In contrast, reinforcement learning-based methods reward subjective human ratings, so the generated reports better capture disease and anatomical structures.
Also, each modality such as CT, MRI, PET, etc. has a specific clinical context, as each modality has a clinically distinct imaging purpose. Generating reports from each modality, therefore, presents unique challenges, as each modality must be evaluated on a case-by-case basis.
the others
Here are some other applications of Transformer. In recent years, Transformer has been remarkably successful in regression tasks to predict the relative risk of death from cancer. A method called MCAT has been proposed by Chen et al. to predict survival based on multimodal information of genetic information and biopsy specimens.
Other proposals include PubMedCLIP (contrastive language-image pre-training) for learning images and captions from PubMed articles and 3DMPT for analyzing 3D medical data.
Future Direction
In this section, we describe the future of ViT in medical imaging.
prior learning
ViT does not have an inductive bias to model local visual features, so it needs to be pre-trained on large datasets. However, collecting large (note: tens to hundreds of millions of images are considered large in the context of Transformer) datasets in the medical field is difficult and can be a limitation. Many reports use pre-training on ImageNet, but natural and medical images have different characteristics. Below is a summary of the characteristics of CNN and ViT.
- When initialized with random weights, CNN outperforms ViT in medical image classification tasks.
- CNN and ViT can also benefit greatly from ImageNet initialization in medical image classification. In particular, ViT seems to bridge the gap between medical and natural images by transition learning.
- CNN and ViT perform well with self-supervised pre-training such as DINO and BYOL, and ViT seems to slightly outperform CNN in this case.

Above is the performance evaluation of ViT pre-trained with medical images, blue is the Swin UNET pre-trained with CT images and orange is the Swin UNET without pre-training. The blue graph compares the die coefficients for Swin UNET pre-trained with CT images and the orange graph compares the die coefficients for Swin UNET without pre-training, indicating that pre-training with medical images is better for Swin UNET.
interpretability
The transformer has been successful in the field of medical imaging but has not yet achieved satisfactory results in terms of interpretability. Although ViT has good performance in medical imaging, it is used in a black box fashion, and although some studies have mentioned interpretability, overall it is still in its infancy. It is not yet possible to explicitly model how the parts of the image interact with each other. Despite the intrinsic suitability of attention for interpretability, the interpretability of medical images is an open problem.
robustness
Advances in adversarial attacks have shown that existing imaging networks are vulnerable to perturbations. Especially in the field of medical imaging related to diagnostics, the huge research budgets inevitably make them vulnerable to attacks. For example, attackers may manipulate medical imaging systems and falsify reports to fraudulently obtain insurance or medical reimbursement.
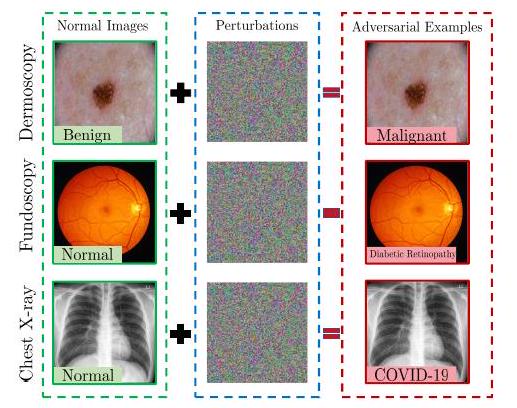
While there are abundant reports on the robustness of CNNs, to the best of our knowledge there are no such reports on ViT. In recent years, several attempts have been made on the attack and robustness of ViT, and when summarized with some differences, ViT is found to be more robust against adversarial attacks than CNNs. As mentioned earlier, due to the different characteristics of natural images and medical images, adversarial attacks also need to be specific to medical images, and the robustness of medical image processing systems with ViT will be evaluated in the future.
Edge AI with ViT
Although ViT has been successful in medical imaging, it is still computationally expensive. This has hindered its deployment in edge computing, where resources are limited. However, there is a high demand for edge computing in the healthcare domain to process, transform, and analyze medical images while preserving patient privacy. While several efforts have been made in recent years to compress Transformer-based models, there is a dire need to design domain-optimized architectures.
Distributed medical image processing solution using ViT
Building robust machine learning models depends on the amount and diversity of training data. Learning reliable and robust models is paradoxically hampered by strict privacy regulations.
Therefore, federated learning (FL) has been proposed for multicenter model building. FL builds a shared model using distributed data from multiple instruments, and each instrument learns with its local data. This allows learning to be performed without sharing patient data, but by sharing parameters with a central model.
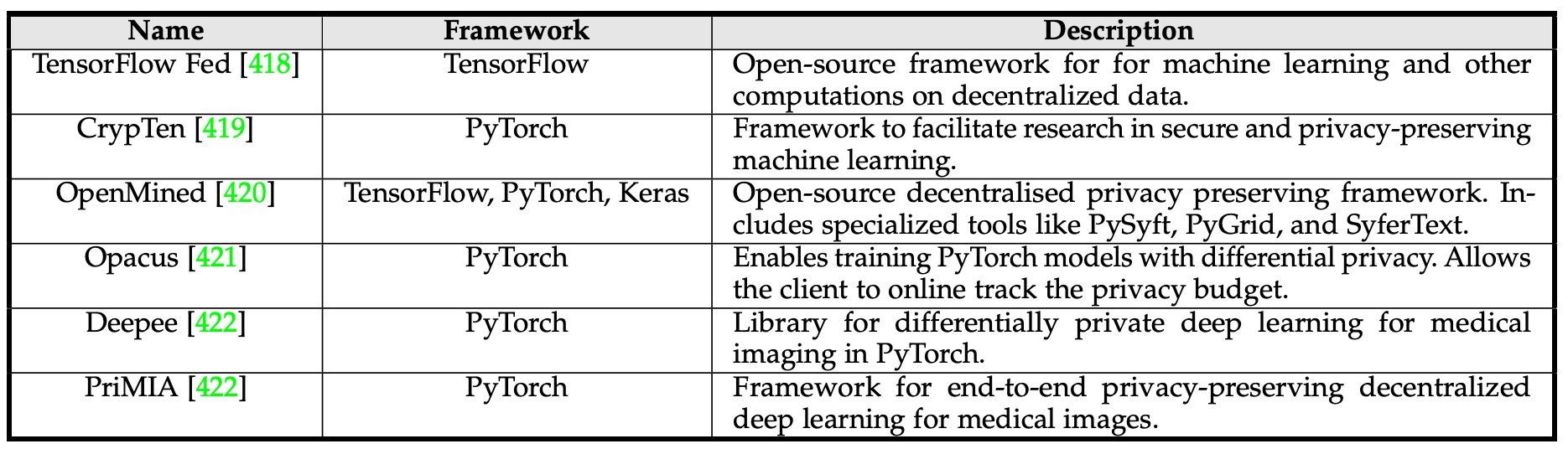
Park et al. proposed a ViT-based system to diagnose COVID-19 in a system based on association learning. However, this model is only a demonstration and will require further clinical validation.
Domain Adaptability and Generalizability
Recent ViT-based medical imaging systems are mainly focused on improving accuracy and lack a mechanism to evaluate generalization capability. Sandhu's work has shown that test errors generally increase in proportion to the difference between the distribution of training and test data. In the context of medical imaging, such distributional discrepancies can be caused by the following factors.
- Images obtained at different facilities and with different equipment
- Diseases not in the training data are included in the test data
- Blurred images with low contrast
On the other hand, however, it has been shown that ViT pre-trained on large-scale data performs well on different modalities. However, most of those models are trained on toy data such as CIFAR-10 and CIFAR100 and are not able to handle complex patterns and local features such as medical images. 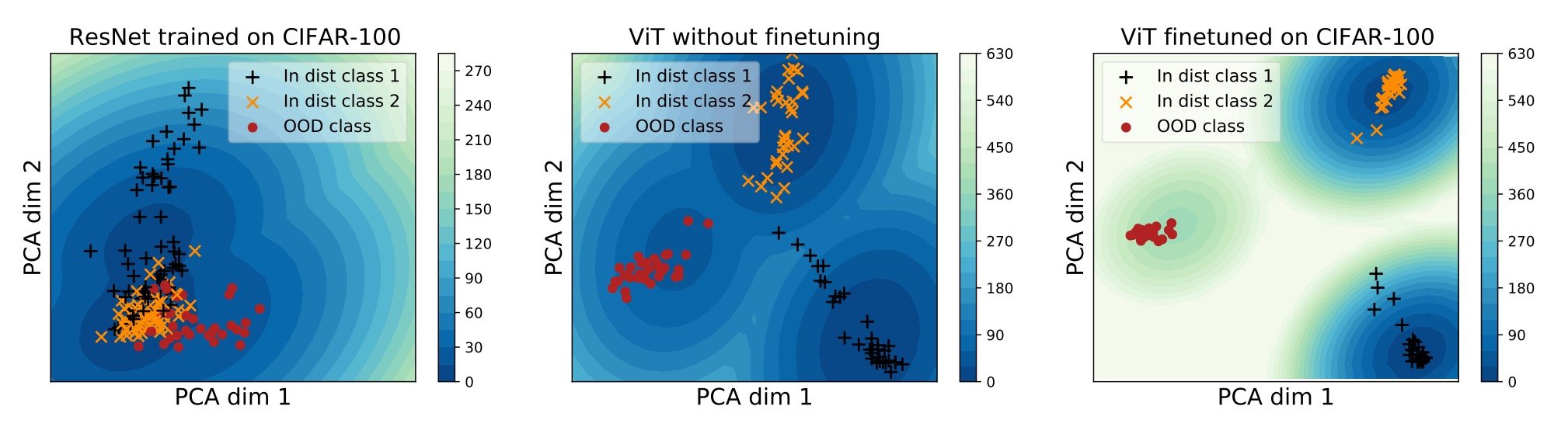
Above is a PCA projection of the embedding vector space of the three models in two dimensions. The ResNet on the left cannot separate the three (yellow, black, and red), but the ViT in the center can classify the classes (but some of the blacks are close to yellow). So, fine tuning ViT with CIFAR-100 can classify them even better.
Discussion and Conclusion
This paper has shown that ViT has penetrated all areas of the medical imaging field. To support this rapid development, it is expected that workshops on CV and medical imaging will be held at conferences and special issues will be published in journals.
With the success of Transformer in the background, this paper provided a comprehensive review of Transformer for classification, detection, segmentation, reconstruction, synthesis, registration, report generation, and other tasks in medical imaging. There is still a lot of exploration left to be done on Transformer in medical imaging and we hope that this paper can provide a better roadmap for ongoing research.
Categories related to this article




![DrHouse] Diagnostic](https://aisholar.s3.ap-northeast-1.amazonaws.com/media/February2025/drhouse-520x300.png)
![SA-FedLoRA] Communic](https://aisholar.s3.ap-northeast-1.amazonaws.com/media/October2024/sa-fedlora-520x300.png)

![[SpliceBERT] A BERT](https://aisholar.s3.ap-northeast-1.amazonaws.com/media/July2024/splicebert-520x300.png)
![[IGModel] Methodolog](https://aisholar.s3.ap-northeast-1.amazonaws.com/media/July2024/igmodel-520x300.png)